Working with Files and Directories
Overview
Teaching: 30 min
Exercises: 15 minQuestions
How can I view and search file contents?
How can I create, copy and delete files and directories?
How can I repeat recently used commands?
Objectives
View, search within, copy, move, and rename files. Create new directories.
Use wildcards (
*) to perform operations on multiple files.Use the
historycommand to view and repeat recently used commands.
Working with Files
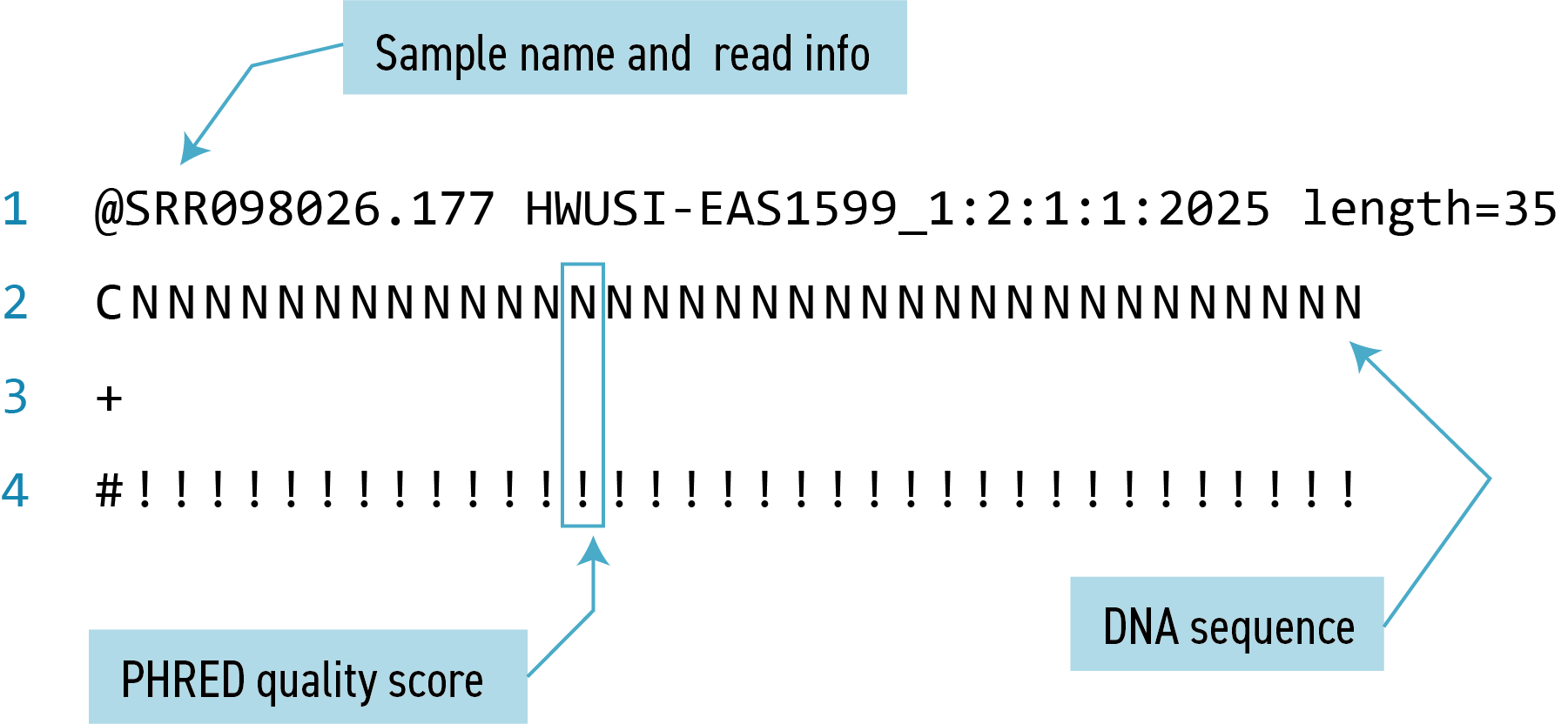
Our data set: FASTQ files
Now that we know how to navigate around our directory structure, let’s
start working with our sequencing files. We did a sequencing experiment and
have two results files, which are stored in our untrimmed_fastq directory.
Wildcards
Navigate to your untrimmed_fastq directory:
$ cd ~/shell_data/untrimmed_fastq
We are interested in looking at the FASTQ files in this directory. We can list all files with the .fastq extension using the command:
$ ls *.fastq
SRR097977.fastq SRR098026.fastq
The * character is a special type of character called a wildcard, which can be used to represent any number of any type of character.
Thus, *.fastq matches every file that ends with .fastq.
This command:
$ ls *977.fastq
SRR097977.fastq
lists only the file that ends with 977.fastq.
This command:
$ ls /usr/bin/*.sh
/usr/bin/amuFormat.sh /usr/bin/gettext.sh /usr/bin/gvmap.sh
Lists every file in /usr/bin that ends in the characters .sh.
Note that the output displays full paths to files, since
each result starts with /.
Exercise
What command would you use for each of the following tasks? Start from your current directory using a single
lscommand for each:
- List all of the files in
/usr/binthat start with the letter ‘c’.- List all of the files in
/usr/binthat contain the letter ‘a’.- List all of the files in
/usr/binthat end with the letter ‘o’.- List all of the files in
/usr/binthat contain the letter ‘a’ or the letter ‘c’.Bonus: What would the output look like if a wildcard could not be matched? Try listing all files that start with ‘missing’.
Hint: Question 4 requires a Unix wildcard that we haven’t talked about yet. Try searching the internet for information about Unix wildcards to find what you need to solve the bonus problem.
Share your answers in the forum.
Solution
ls /usr/bin/c*ls /usr/bin/*a*ls /usr/bin/*ols /usr/bin/*[ac]*Bonus:
ls: cannot access 'missing*': No such file or directory
Command History
If you want to repeat a command that you’ve run recently, you can access previous commands using the up arrow on your keyboard to go back to the most recent command. Likewise, the down arrow takes you forward in the command history.
A few more useful shortcuts:
- Ctrl+C will cancel the command you are writing, and give you a fresh prompt.
- Ctrl+R will do a reverse-search through your command history. This is very useful.
- Ctrl+L or the
clearcommand will clear your screen.
You can also review your recent commands with the history command, by entering:
$ history
to see a numbered list of recent commands. You can reuse one of these commands directly by referring to the number of that command.
For example, if your history looked like this:
259 ls *
260 ls /usr/bin/*.sh
261 ls *R1*fastq
then you could repeat command #260 by entering:
$ !260
Type ! (exclamation point) and then the number of the command from your history.
You will be glad you learned this when you need to re-run very complicated commands.
For more information on advanced usage of history, read section 9.3 of
Bash manual.
Examining Files
We now know how to switch directories, run programs, and look at the contents of directories, but how do we look at the contents of files?
One way to examine a file is to print out all of the
contents using the program cat.
Enter the following command from within the untrimmed_fastq directory:
$ cat SRR098026.fastq
This will print out all of the contents of the SRR098026.fastq to the screen.
Reminder
The contents might look a bit confusing. As these are
.fastqfiles, they use the FASTQ format we have discussed previously. Here is a reminder to keep in mind as we continue to examine these files:
Exercise
- Print out the contents of the
~/shell_data/untrimmed_fastq/SRR097977.fastqfile. What is the last line of the file?- From your home directory, and without changing directories, use one short command to print the contents of all of the files in the
~/shell_data/untrimmed_fastqdirectory.Share your answers on the forum.
Solution
- The last line of the file is
C:CCC::CCCCCCCC<8?6A:C28C<608'&&&,'$.cat ~/shell_data/untrimmed_fastq/*
cat is a terrific program, but when the file is really big, it can
be annoying to use. The program, less, is useful for this
case. less opens the file as read only, and lets you navigate through it. The navigation commands
are identical to the man program.
Enter the following command:
$ less SRR097977.fastq
Some navigation commands in less:
| key | action |
|---|---|
| Space | to go forward |
| b | to go backward |
| g | to go to the beginning |
| G | to go to the end |
| q | to quit |
less also gives you a way of searching through files. Use the
“/” key to begin a search. Enter the word you would like
to search for and press enter. The screen will jump to the next location where
that word is found.
Shortcut: If you hit “/” then “enter”, less will repeat
the previous search. less searches from the current location and
works its way forward. Scroll up a couple lines on your terminal to verify
you are at the beginning of the file. Note, if you are at the end of the file and search
for the sequence “CAA”, less will not find it. You either need to go to the
beginning of the file (by typing g) and search again using / or you
can use ? to search backwards in the same way you used / previously.
For instance, let’s search forward for the sequence TTTTT in our file.
You can see that we go right to that sequence, what it looks like,
and where it is in the file. If you continue to type / and hit return, you will move
forward to the next instance of this sequence motif. If you instead type ? and hit
return, you will search backwards and move up the file to previous examples of this motif.
Exercise
What are the next three nucleotides (characters) after the first instance of the sequence quoted above (
TTTTT)?Share your answer on the forum to see if it matches everyone else’s!
Solution
CAC
Remember, the man program actually uses less internally and
therefore uses the same commands, so you can search documentation
using “/” as well!
There’s another way that we can look at files, and in this case, just look at part of them. This can be particularly useful if we just want to see the beginning or end of the file, or see how it’s formatted.
The commands are head and tail and they let you look at
the beginning and end of a file, respectively.
$ head SRR098026.fastq
@SRR098026.1 HWUSI-EAS1599_1:2:1:0:968 length=35
NNNNNNNNNNNNNNNNCNNNNNNNNNNNNNNNNNN
+SRR098026.1 HWUSI-EAS1599_1:2:1:0:968 length=35
!!!!!!!!!!!!!!!!#!!!!!!!!!!!!!!!!!!
@SRR098026.2 HWUSI-EAS1599_1:2:1:0:312 length=35
NNNNNNNNNNNNNNNNANNNNNNNNNNNNNNNNNN
+SRR098026.2 HWUSI-EAS1599_1:2:1:0:312 length=35
!!!!!!!!!!!!!!!!#!!!!!!!!!!!!!!!!!!
@SRR098026.3 HWUSI-EAS1599_1:2:1:0:570 length=35
NNNNNNNNNNNNNNNNANNNNNNNNNNNNNNNNNN
$ tail SRR098026.fastq
+SRR098026.247 HWUSI-EAS1599_1:2:1:2:1311 length=35
#!##!#################!!!!!!!######
@SRR098026.248 HWUSI-EAS1599_1:2:1:2:118 length=35
GNTGNGGTCATCATACGCGCCCNNNNNNNGGCATG
+SRR098026.248 HWUSI-EAS1599_1:2:1:2:118 length=35
B!;?!A=5922:##########!!!!!!!######
@SRR098026.249 HWUSI-EAS1599_1:2:1:2:1057 length=35
CNCTNTATGCGTACGGCAGTGANNNNNNNGGAGAT
+SRR098026.249 HWUSI-EAS1599_1:2:1:2:1057 length=35
A!@B!BBB@ABAB#########!!!!!!!######
The -n option to either of these commands can be used to print the
first or last n lines of a file.
$ head -n 1 SRR098026.fastq
@SRR098026.1 HWUSI-EAS1599_1:2:1:0:968 length=35
$ tail -n 1 SRR098026.fastq
A!@B!BBB@ABAB#########!!!!!!!######
You will learn more about how to interpret these outputs on day three of the Cloud-SPAN Genomics module.
Creating, moving, copying, and removing
Now we can move around in the file structure, look at files, and search files. But what if we want to copy files or move them around or get rid of them? Most of the time, you can do these sorts of file manipulations without the command line, but there will be some cases (like when you’re working with a remote computer like we are for this lesson) where it will be impossible. You’ll also find that you may be working with hundreds of files and want to do similar manipulations to all of those files. In cases like this, it’s much faster to do these operations at the command line.
Copying Files
When working with computational data, it’s important to keep a safe copy of that data that can’t be accidentally overwritten or deleted. For this lesson, our raw data is our FASTQ files.
First, let’s make a copy of one of our FASTQ files using the cp command.
Navigate to the shell_data/untrimmed_fastq directory and enter:
$ cp SRR098026.fastq SRR098026-copy.fastq
$ ls -F
SRR097977.fastq SRR098026-copy.fastq SRR098026.fastq
We now have two copies of the SRR098026.fastq file, one of them named SRR098026-copy.fastq. We’ll move this file to a new directory
called backup where we’ll store our backup data files.
Creating Directories
The mkdir command is used to make a directory. Enter mkdir
followed by a space, then the directory name you want to create:
$ mkdir backup
Moving / Renaming
We can now move our backup file to this directory. We can
move files around using the command mv:
$ mv SRR098026-copy.fastq backup
$ ls backup
SRR098026-copy.fastq
The mv command is also how you rename files. Let’s rename this file to make it clear that this is a backup:
$ cd backup
$ mv SRR098026-copy.fastq SRR098026-backup.fastq
$ ls
SRR098026-backup.fastq
Removing
You can delete or remove files with the rm command:
$ rm SRR098026-backup.fastq
Important: The rm command permanently removes the file. Be careful with this command. It doesn’t just nicely put the files in the Trash. They’re really gone.
By default, rm will not delete directories. You can tell rm to
delete a directory using the -r (recursive) option. Let’s delete the backup directory
we just made.
Enter the following command:
$ cd ..
$ rm -r backup
This will delete not only the directory, but all files within the directory.
Exercise
Starting in the
shell_data/untrimmed_fastq/directory, do the following:
- Make sure that you have deleted your backup directory and all files it contains.
- Create a backup of each of your FASTQ files using
cp. (Note: You’ll need to do this individually for each of the two FASTQ files. We haven’t learned yet how to do this with a wildcard.)- Use a wildcard to move all of your backup files to a new backup directory.
- It doesn’t make sense to keep our backup directory inside the directory it is backing up. What if we accidentally delete the
untrimmed_fastqdirectory? To fix this, move your new backup directory out ofuntrimmed_fastqand into the parent folder,shell_data.Solution
rm -r backupcp SRR098026.fastq SRR098026-backup.fastqandcp SRR097977.fastq SRR097977-backup.fastqmkdir backupandmv *-backup.fastq backupmv backup ..ormv backup ~/shell_data/(note that you do not need to use the -r flag to move directories like you do when deleting them)It’s always a good idea to check your work. Move to the
shell_datafolder withcd ..and then list the contents ofbackupwithls -l backup. You should see something like:-rw-r--r-- 1 csuser csuser 47552 Nov 15 23:06 SRR097977-backup.fastq -rw-r--r-- 1 csuser csuser 43332 Nov 15 23:06 SRR098026-backup.fastq
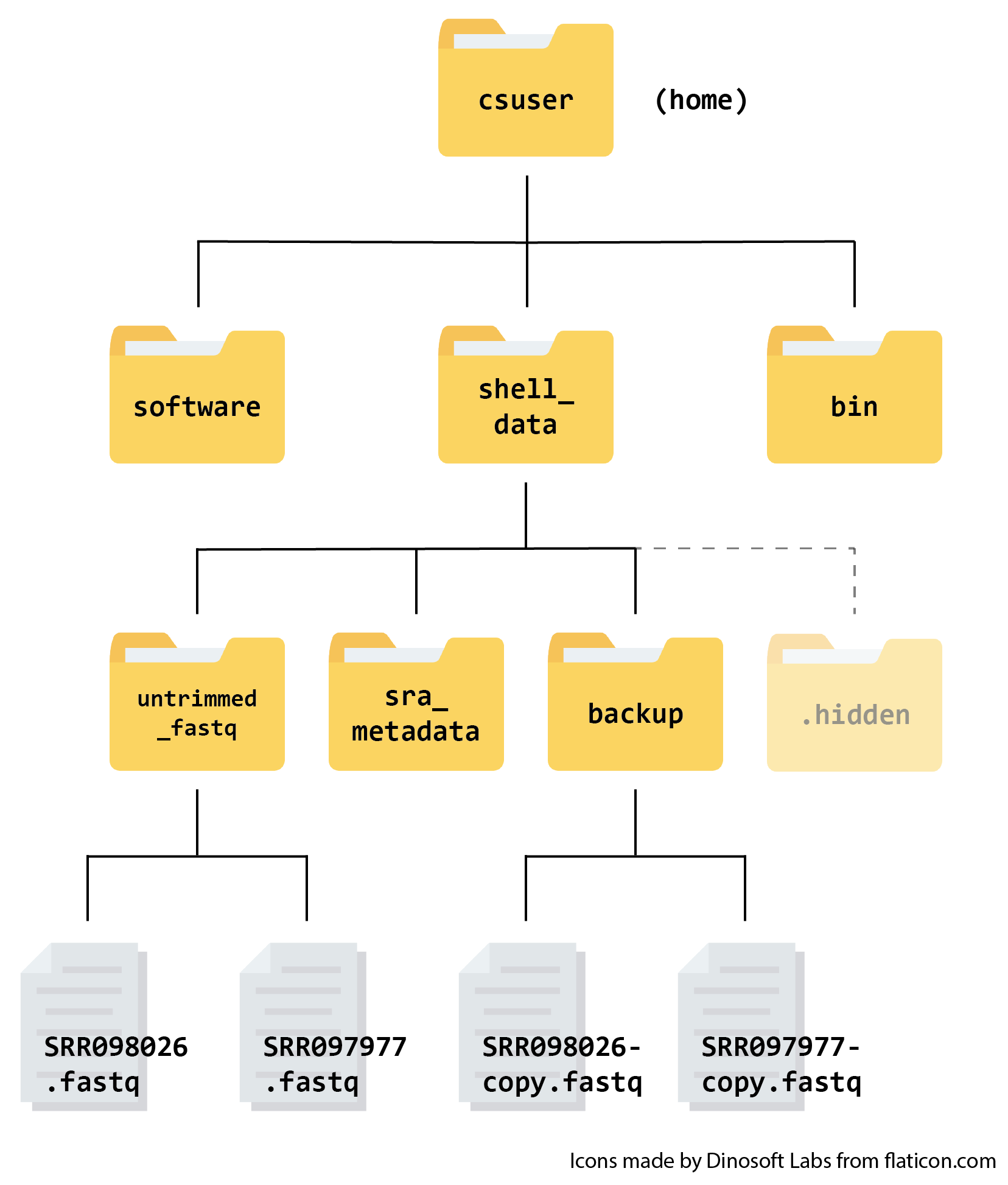
Here is what your file structure should look like at the end of this episode:
Key Points
You can view file contents using
less,cat,headortail.The commands
cp,mv, andmkdirare useful for manipulating existing files and creating new directories.The
historycommand and the up arrow on your keyboard can be used to repeat recently used commands.